[ad_1]
Summary: A new study of gene expression in the hippocampus has unveiled two new genes potentially implicated in Alzheimer’s disease.
Source: PLOS
A research team led by Chunshui Yu and Mulin Jun Li of Tianjin Medical University has discovered two new genes potentially involved in Alzheimer’s disease. They identified them by exploring which genes were turned on and off in the hippocampus of people who suffered from the disease.
The team’s new findings are published February 25th in PLOS Genetics.
Alzheimer’s disease is a neurodegenerative disorder that involves worsening dementia and the formation of protein plaques and tangles in the brain. The hippocampus, part of the brain involved in memory, is one of the first regions to sustain damage.
To better understand which genes contribute to the progression of this heritable disease, the researchers identified genes expressed at higher or lower levels in the hippocampus of people with Alzheimer’s disease compared to healthy brains.

They identified 24 Alzheimer’s-related genes that appear to have an effect through the hippocampus, using previous genomic and hippocampus gene expression data. Many genes were already known to contribute to the disease, such as APOE, but two were unknown, PTPN9 and PCDHA4. Additionally, several are involved in biological process related to Alzheimer’s disease, such as plaque formation and cell death.
The research team further validated their findings by comparing gene expression for the two dozen genes to images of the individuals’ brains. In Alzheimer’s disease, damage and loss of neurons causes the hippocampus to shrink, which can be measured through medical imaging. The researchers established that expression of two of the genes is related to the size of the hippocampus and a diagnosis of Alzheimer’s disease.
Overall, the new findings improve our understanding of the genetic and cellular mechanisms that cause Alzheimer’s disease. The next step will be to investigate the roles of the two novel genes and how they contribute to this devastating disease.
The authors add, “The study identifies two novel genes associated with Alzheimer’s disease in the context of hippocampal tissue and reveals candidate hippocampus-mediated neurobiological pathways from gene expression to Alzheimer’s disease.”
Funding: This work was supported by the National Key Research and Development Program of China (2018YFC1314300) to CY, the National Natural Science Foundation of China to CY (82030053, 81425013), the National Natural Science Foundation of China to MJL (31871327), the National Natural Science Foundation of China to JX (82001797), the National Natural Science Foundation of China to HL (81701668) and the Tianjin Key Technology R&D Program (17ZXMFSY00090) to CY. The authors report no biomedical financial interests or potential conflicts of interest. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
About this genetics and Alzheimer’s disease research news
Source: PLOS
Contact: Press Office – PLOS
Image: The image is credited to Liu N, et al., 2021, PLOS Genetics
Original Research: Open access.
“Hippocampal transcriptome-wide association study and neurobiological pathway analysis for Alzheimer’s disease” by Liu N, Xu J, Liu H, Zhang S, Li M, Zhou Y, et al. PLOS Genetics
Abstract
Hippocampal transcriptome-wide association study and neurobiological pathway analysis for Alzheimer’s disease
Genome-wide association studies (GWASs) have identified multiple susceptibility loci for Alzheimer’s disease (AD), which is characterized by early and progressive damage to the hippocampus. However, the association of hippocampal gene expression with AD and the underlying neurobiological pathways remain largely unknown.
Based on the genomic and transcriptomic data of 111 hippocampal samples and the summary data of two large-scale meta-analyses of GWASs, a transcriptome-wide association study (TWAS) was performed to identify genes with significant associations between hippocampal expression and AD. We identified 54 significantly associated genes using an AD-GWAS meta-analysis of 455,258 individuals; 36 of the genes were confirmed in another AD-GWAS meta-analysis of 63,926 individuals.
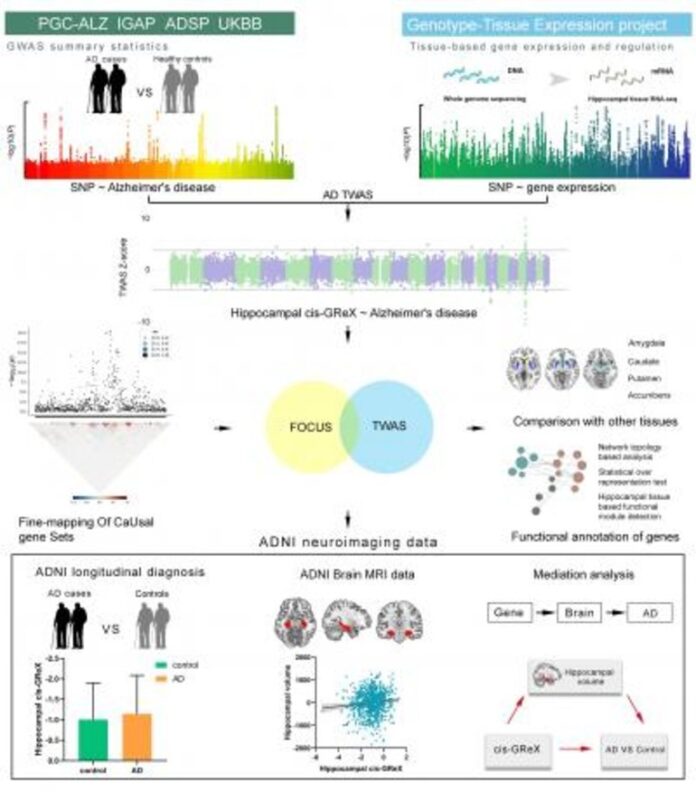
Fine-mapping models further prioritized 24 AD-related genes whose effects on AD were mediated by hippocampal expression, including APOE and two novel genes (PTPN9 and PCDHA4). These genes are functionally related to amyloid-beta formation, phosphorylation/dephosphorylation, neuronal apoptosis, neurogenesis and telomerase-related processes. By integrating the predicted hippocampal expression and neuroimaging data, we found that the hippocampal expression of QPCTL and ERCC2 showed significant difference between AD patients and cognitively normal elderly individuals as well as correlated with hippocampal volume. Mediation analysis further demonstrated that hippocampal volume mediated the effect of hippocampal gene expression (QPCTL and ERCC2) on AD.
This study identifies two novel genes associated with AD by integrating hippocampal gene expression and genome-wide association data and reveals candidate hippocampus-mediated neurobiological pathways from gene expression to AD.
[ad_2]
Source link